-Search query
-Search result
Showing 1 - 50 of 116 items for (author: jiang & yl)

EMDB-44635: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM
Method: single particle / : O'Brien ES, Wang H, Kaavya Krishna K, Zhang C, Kobilka BK

PDB-9bjk: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM
Method: single particle / : O'Brien ES, Wang H, Kaavya Krishna K, Zhang C, Kobilka BK

EMDB-39064: 
Structure of NET-Maprotiline in outward-open state
Method: single particle / : Zhang H, Xu EH, Jiang Y
EMDB-39065: 
Structure of NET-Nefopam in outward-open state
Method: single particle / : Zhang H, Xu EH, Jiang Y

EMDB-39066: 
Structure of NET-nomifensine in outward-open state
Method: single particle / : Zhang H, Xu EH, Jiang Y

EMDB-39067: 
structure of NET-Atomoxetine in outward-open state
Method: single particle / : Zhang H, Xu EH, Jiang Y

EMDB-39068: 
Structure of NET-Amitriptyline in outward-open state
Method: single particle / : Zhang H, Xu EH, Jiang Y

EMDB-39069: 
Structure of Apo human norepinephrine transporter NET
Method: single particle / : Zhang H, Xu HE, Jiang Y

EMDB-39070: 
Structure of NET-NE in Occluded state
Method: single particle / : Zhang H, Xu HE, Jiang Y

EMDB-39533: 
Structure of NET-Nisoxetine in outward-open state
Method: single particle / : Zhang H, Xu EH, Jiang Y

EMDB-37727: 
Cryo-ET structure of RuBisCO from 3.9 angstroms Synechococcus elongatus PCC 7942
Method: subtomogram averaging / : Kong WW, Jiang YL, Zhou CZ

EMDB-37728: 
Cryo-ET map of RuBisCO at 4.4 angstroms from Synechococcus elongatus PCC 7942 beta-carboxysome
Method: subtomogram averaging / : Kong WW, Jiang YL, Zhou CZ

EMDB-37729: 
Cryo-ET map of RuBisCO-SSUL at 5.9 angstroms from Synechococcus elongatus PCC 7942 beta-carboxysome
Method: subtomogram averaging / : Kong WW, Jiang YL, Zhou CZ

EMDB-37730: 
Cryo-ET map of RuBisCO at the outermost layer that is loosely attached to the shell of Synechococcus elongatus PCC 7942 beta-carboxysome
Method: subtomogram averaging / : Kong WW, Jiang YL, Zhou CZ

EMDB-37731: 
Cryo-ET map of RuBisCO at the outermost layer that is tightly attached to the shell of Synechococcus elongatus PCC 7942 beta-carboxysome
Method: subtomogram averaging / : Kong WW, Jiang YL, Zhou CZ

EMDB-37375: 
Allosteric regulation of nitrate transporter NRT via the signalling protein PII
Method: single particle / : Li B, Zhou CZ, Chen YX, Jiang YL

EMDB-37644: 
Cryo-EM structure of cyanobacterial nitrate/nitrite transporter NrtBCD in complex with signalling protein PII
Method: single particle / : Li B, Zhou CZ, Chen YX, Jiang YL

EMDB-37645: 
Cryo-EM structure of cyanobacterial nitrate/nitrite transporter NrtBCD in complex with nitrate
Method: single particle / : Li B, Zhou CZ, Chen YX, Jiang YL

EMDB-38543: 
Cryo-EM map of the internal RuBisCOs in the alpha-carboxysome from Prochlorococcus MED4
Method: single particle / : Jiang YL, Zhou RQ, Zhou CZ, Zeng QL

EMDB-38544: 
Cryo-EM map of the intact shell of alpha-carboxysome from Prochlorococcus MED4
Method: single particle / : Jiang YL, Zhou RQ, Zhou CZ, Zeng QL

EMDB-37902: 
Cryo-EM structure of the alpha-carboxysome shell vertex from Prochlorococcus MED4
Method: single particle / : Jiang YL, Zhou RQ, Zhou CZ, Zeng QL

EMDB-37903: 
Cryo-EM map of the intact alpha-carboxysome from Prochlorococcus MED4
Method: single particle / : Jiang YL, Zhou RQ, Zhou CZ, Zeng QL

EMDB-29944: 
Cryo-EM Structure of the Prostaglandin E2 Receptor 4 Coupled to G Protein
Method: single particle / : Shenming H, Mengyao X, Lei L, Jiangqian M, Sheng C, Jinpeng S
EMDB-40976: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to two mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

EMDB-40977: 
Cryo-EM structure of mink variant Y453F trimeric spike protein
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B
EMDB-40978: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors at downRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

EMDB-40979: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at upRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

EMDB-40980: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at downRBD conformation.
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B
EMDB-41143: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Liang B

PDB-8t20: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to two mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t21: 
Cryo-EM structure of mink variant Y453F trimeric spike protein
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t22: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors at downRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t23: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at upRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t25: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at downRBD conformation.
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8taz: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Liang B

EMDB-34475: 
Cryo-EM structure of the full transcription activation complex NtcA-NtcB-TAC
Method: single particle / : Han SJ, Jiang YL, You LL, Shen LQ, Wu XX, Yang F, Kong WW, Chen ZP, Zhang Y, Zhou CZ

EMDB-34476: 
Cryo-EM structure of the transcription activation complex NtcA-TAC
Method: single particle / : Han SJ, Jiang YL, You LL, Shen LQ, Wu XX, Yang F, Kong WW, Chen ZP, Zhang Y, Zhou CZ

EMDB-34477: 
Cryo-EM structure of the full transcription activation complex NtcA-NtcB-TAC focusing on NtcA and NtcB binding sites
Method: single particle / : Han SJ, Jiang YL, You LL, Shen LQ, Kong WW, Zhang Y, Zhou CZ

EMDB-29735: 
Structure of nucleosome-bound Sirtuin 6 deacetylase
Method: single particle / : Chio US, Rechiche O, Bryll AR, Zhu J, Feldman JL, Peterson CL, Tan S, Armache JP

PDB-8g57: 
Structure of nucleosome-bound Sirtuin 6 deacetylase
Method: single particle / : Chio US, Rechiche O, Bryll AR, Zhu J, Feldman JL, Peterson CL, Tan S, Armache JP
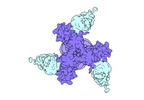
EMDB-27706: 
Vaccine elicited Antibody MU89 bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
Method: single particle / : Stalls V, Acharya P
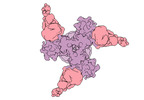
EMDB-27776: 
Vaccine elicited Antibody MU89+S27Y bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
Method: single particle / : Stalls V, Acharya P

PDB-8dto: 
Vaccine elicited Antibody MU89 bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
Method: single particle / : Stalls V, Acharya P

PDB-8dy6: 
Vaccine elicited Antibody MU89+S27Y bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
Method: single particle / : Stalls V, Acharya P

EMDB-33320: 
Cryo-EM map of hMCM-DH R195A/L209G mutant
Method: single particle / : Li J, Dong JQ, Dang SY, Zhai YL

EMDB-34678: 
Cyanophage Pam3 neck
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-33799: 
Cyanophage Pam3 fiber
Method: single particle / : Yang F, Jiang YL, Zhou CZ
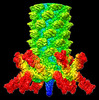
EMDB-33802: 
Cyanophage Pam3 baseplate proteins
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-34679: 
Cyanophage Pam3 portal-adaptor
Method: single particle / : Yang F, Jiang YL, Zhou CZ

EMDB-34680: 
Cyanophage Pam3 capsid asymmetric unit
Method: single particle / : Yang F, Jiang YL, Zhou CZ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
